‘Instagram’ of the immune system predicts response to anti-PD-1 immunotherapy in melanoma
Since the beginning of my scientific career, reaching back to my graduate studies, I was always interested in combining cell biology with conventional flow cytometry, and I followed the rapid development of the flow cytometry field as it went from being able to monitor four parameters all the way up to 16 parameters. In 2014, as an independent fellow with a focus on translational immunology in the University Research Priority Program (URPP) in Zurich, Switzerland and while still deciding on whether to pursue a career in academic science or industry, I had the opportunity to join the laboratory of Burkhard Becher. At that time Burkhard was returning from a sabbatical in the laboratory of Evan Newell at the A*Star Institute, Singapore, where he got insights into the new technology of mass cytometry (MC). MC uses rare metal-labeled probes instead of the more common fluorochrome-labeled probes, thus avoiding spectral overlap from conventional flow, tissue auto-fluorescence and being able to potentially detect up to 100 markers per cell (although, due to practical restrictions, about 50 markers are more realistic to date).
Whereas Burkhard started to use MC in preclinical models1 and wanted to expand his MC research into autoimmunity, I could use MC for translational research in cancer patients. In this context, I was very fortunate to know Mitchell P. Levesque who, together with Reinhard Dummer, just finished one of the first clinical trials on the efficacy of anti-programmed cell death protein 1 (PD-1) in metastatic melanoma patients. It was evident that while immune checkpoint blockade had revolutionized cancer therapy and resulted in a dramatic increase in progression-free survival, a large proportion of patients did not show durable response or relapsed at a later time point. Therefore, we had the idea to employ MC combined with bioinformatics for the in-depth characterization of the immune cell subsets for biomarker discovery in order to understand whether immune parameters could be used to discriminate between responders and non-responders before treatment initiation.
Using bioinformatics would enable us to simplify the complex data set into one visual picture to create a so-called ‘Instagram’ of the immune status of the patient during a specific time point. Early in the project, it became evident that, due to the group number and multiple time points, the off-the-shelf bioinformatic solutions at hand were not sufficient to address our questions. We decided to team up with Mark Robinson at the University of Zurich who, together with his graduate student Malgorzata Nowicka, created a general purpose bioinformatic pipeline to probe the peripheral blood of stage IV melanoma patients before and after 12 weeks of anti-PD-1 immunotherapy2. This was new territory and we were extremely excited about the outcomes.
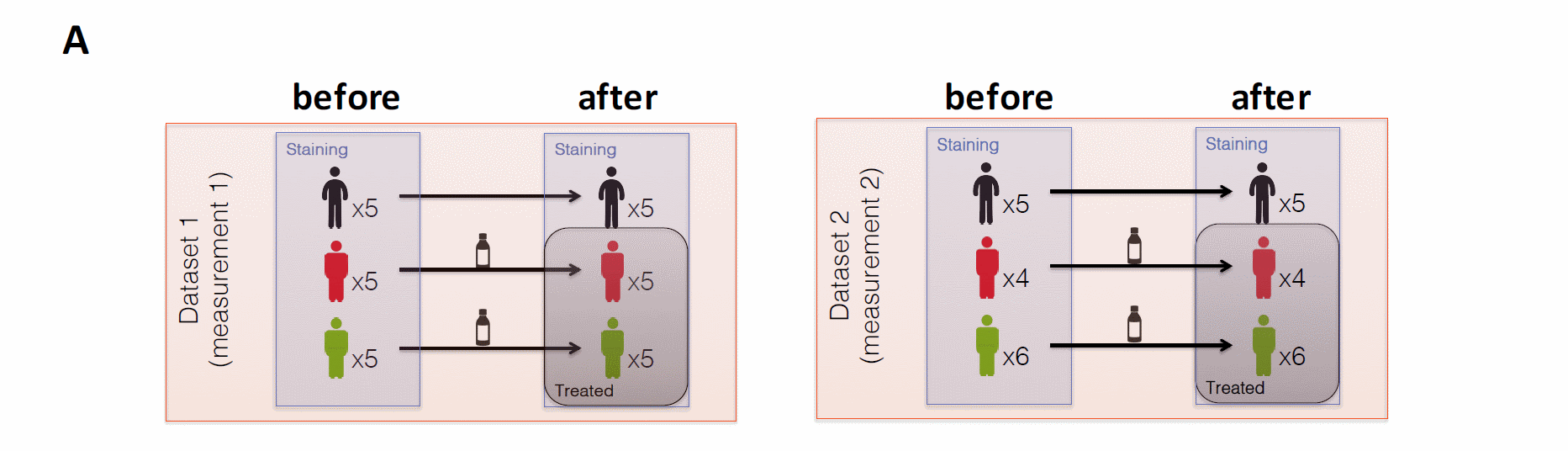
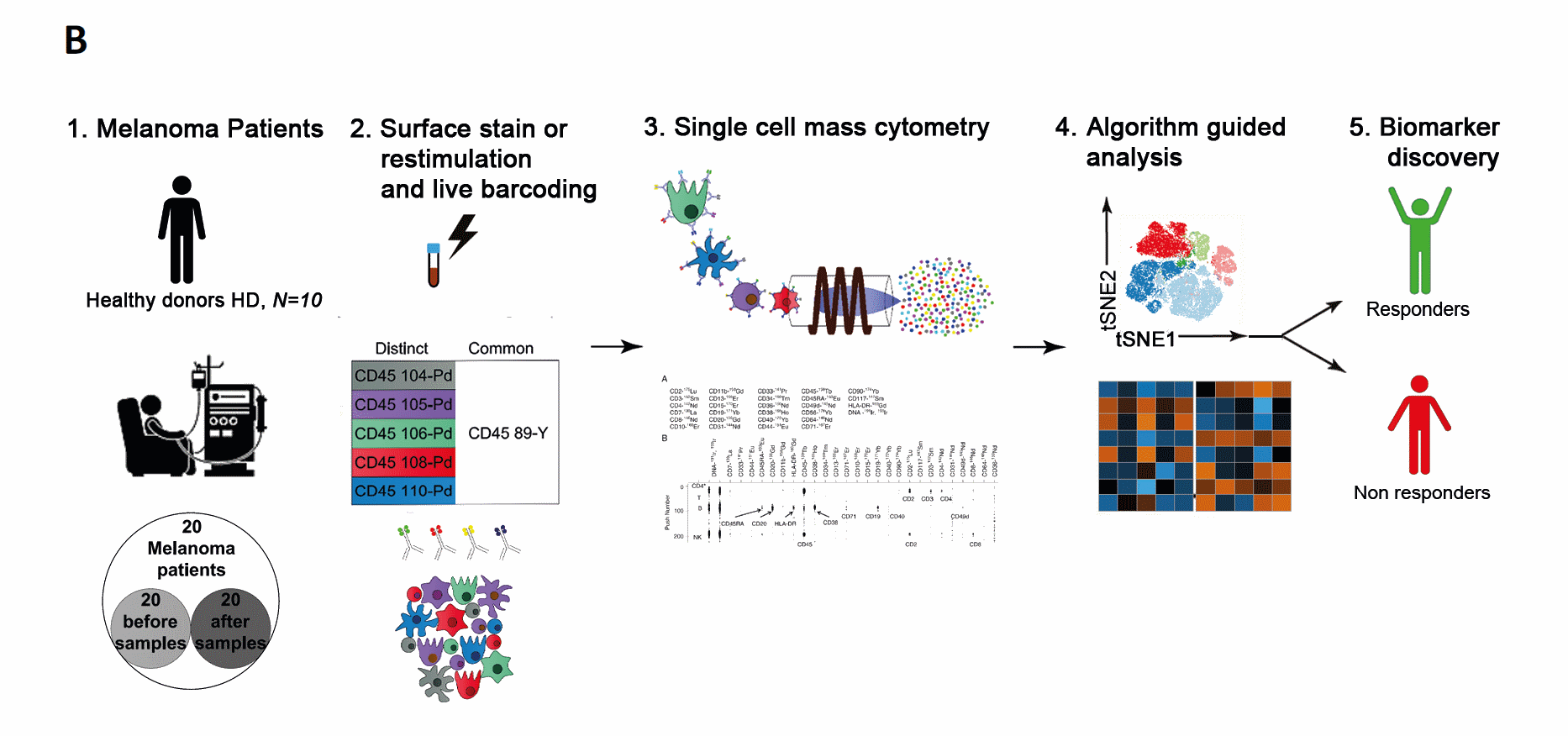
Figure 1. Experimental design for the discovery cohort using CyTOF. Experimental setup for the processing of frozen PBMC from matched samples before and after PD-1 immunotherapy. (A) Total samples n=60 were distributed over 2 datasets. 20 melanoma patients before (R=11 and NR=9) and after (12 weeks treatment) treatment initiation and 10 healthy controls using metal-labeled antibodies and (B) subsequent processing of samples from A for the acquisition by mass cytometry and final bioinformatics analysis. CyTOF, cytometry by time-of-flight; PBMC, peripheral blood mononuclear cells.
As expected based on published results, our first two T cell-centric panels showed a clear treatment response during immunotherapy in the T cell compartment. As we also ran a parallel project with leukemia patients, we decided to probe the myeloid compartment in the peripheral blood of our anti-PD-1 melanoma patients. To our surprise, we discovered that prior to commencing therapy, a strong predictor of progression-free and overall survival in responders to anti-PD-1 immunotherapy was the increased frequency of CD14+CD16–HLA-DRhi monocytes.
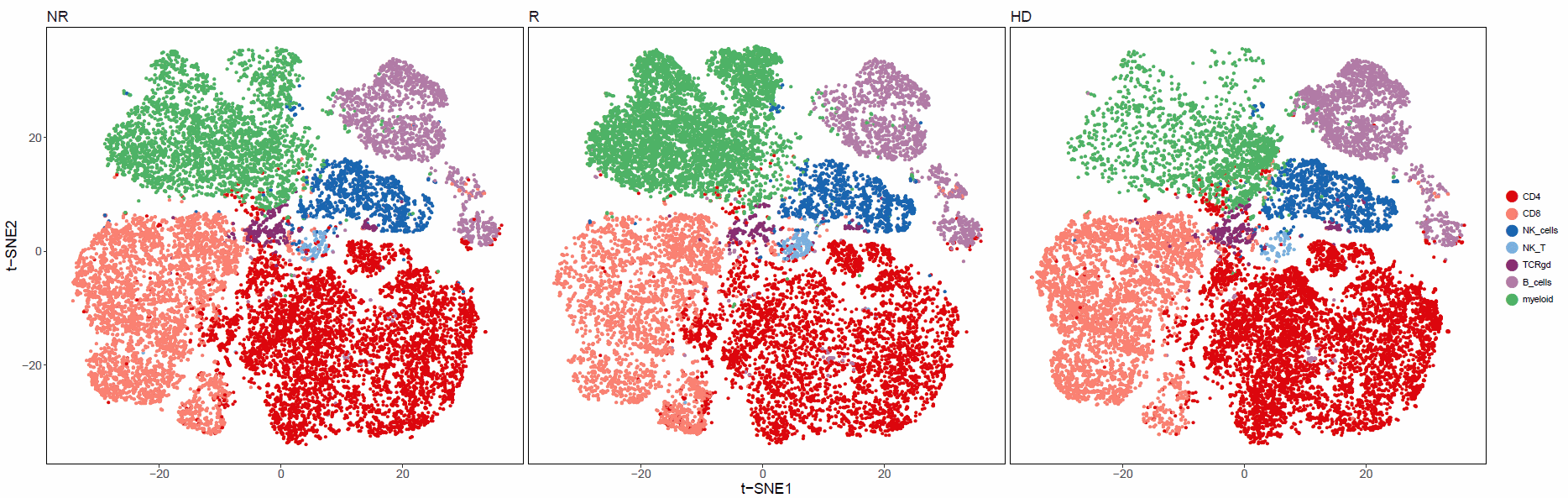
Figure 2. Visualization of patient stratification based on myeloid cell markers and expansion of classical monocytes in responders before anti-PD-1 immunotherapy. Exemple of tSNE visualization of 5,000 merged events from non-responders (NR, n=5), responders (R, n=5), before therapy and healthy donors (HD, n=5), respectively. Cells are colored according to the cluster they were assigned to using the FlowSOM algorithm and manual annotation.
Working on this project gave me the opportunity to experience and incredibly broaden my knowledge of and experience with new technologies. I was lucky to be a member of a highly collaborative team of scientists—reaching from clinicians to wet lab scientists—that made it possible to obtain and report these data3. I am now an Assistant Professor at the Medical University of South Carolina (MUSC), USA. We hope to use MC on both sides of the Atlantic to predict response to other immunotherapeutics and cancers, and to provide tools to inform decision-making when assessing response to anti-PD-1 immunotherapy in the clinic.
OUR PAPER: Krieg, C.*, Nowicka, M., Guglietta, S., Schindler, S., Hartmann, F. J., Weber, L. M., Dummer, R., Robinson, M. D., Levesque, M. P.* & Becher, B.*. High dimensional analysis of the immune landscape during anti-PD-1 immunotherapy of melanoma predicts responsiveness. Nat. Med. doi:10.1038/nm.4466 (*Corresponding authors).
The paper in Nature Medicine is here: http://go.nature.com/2mdJFc6
REFERENCES
1. Becher, B.et al. High-dimensional analysis of the murine myeloid cell system. Nat. Immunol. 15:1181-1189 (2014).
2. Nowicka, M. et al. CyTOF workflow: differential discovery in high-throughput high-dimensional cytometry datasets. F1000Research 6, 748 (2017).
3. Krieg, C. et al. High dimensional analysis of the immune landscape during anti-PD-1 immunotherapy of melanoma predicts responsiveness. Nat. Med. doi:10.1038/nm.4466.
We acknowledge Cinzia Guglietta for the graphical concept of this study.